At the core of S2P Dynamics is a patent-pending deep learning architecture designed specifically for molecular property prediction in drug and radiotracer discovery.
Our approach builds on recent advances in graph neural networks and transformer-style attention, combining them into a unified system tailored for chemical data.
How Scaffold™ works
Click each card to learn more
Unified Molecular Modeling
Unified Molecular Modeling
Our hybrid graph-transformer architecture captures both local atomic interactions and global molecular context in a single unified model. By learning long-range dependencies and higher-level chemical signals simultaneously, the platform uncovers subtle structure–property relationships essential for accurately predicting CNS-relevant outcomes such as blood–brain barrier permeability and ADMET behavior.
Proprietary Feature Fusion
Proprietary Feature Fusion
We integrate deep, learned molecular representations with curated physicochemical descriptors through a proprietary feature-fusion strategy. This approach combines the pattern-recognition power of deep learning with established chemical knowledge, delivering greater robustness, stronger generalization, and consistently high performance across diverse chemical spaces.
Interpretable by Design
Interpretable by Design
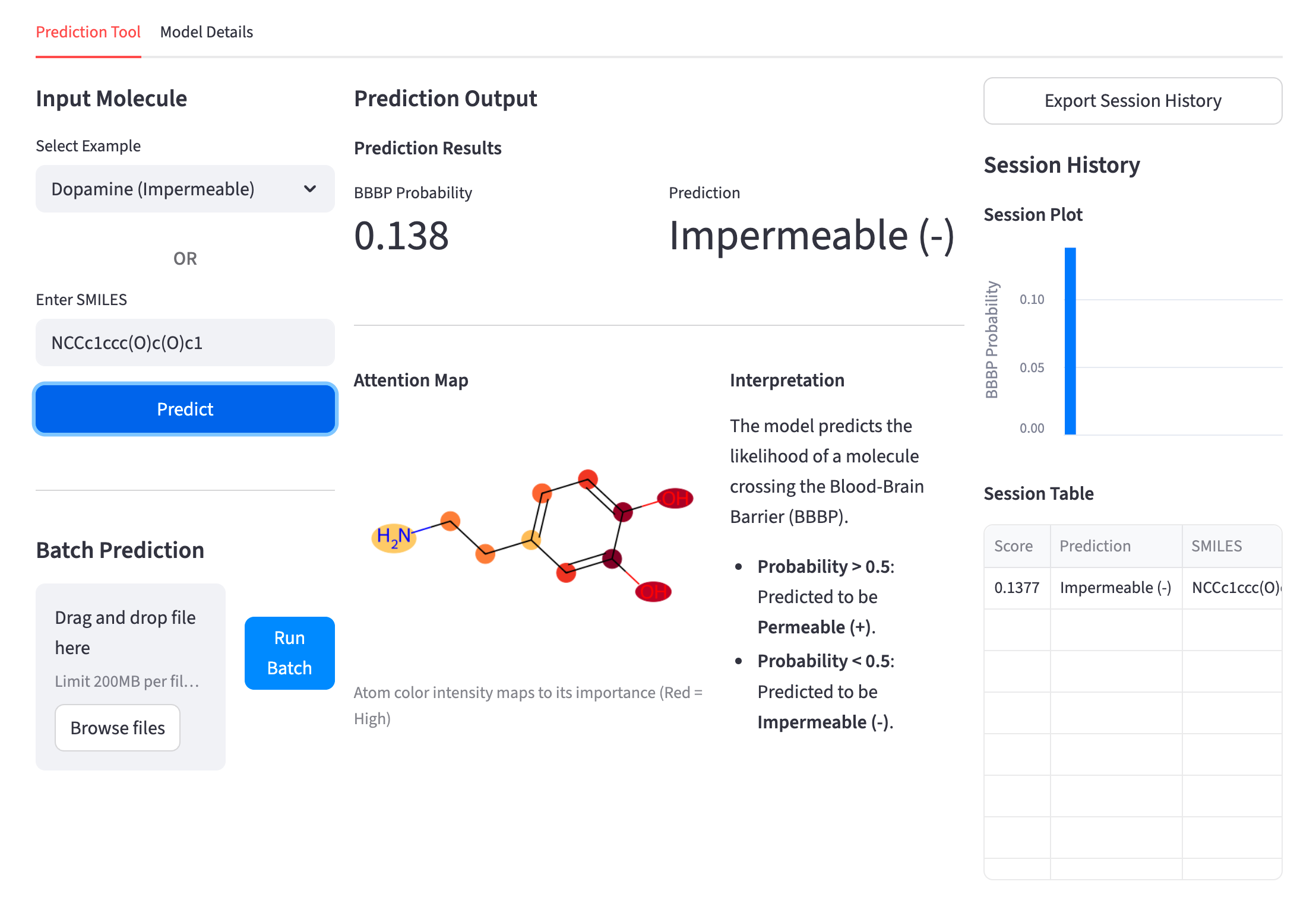
Our models are designed to explain their predictions. Attention mechanisms highlight the molecular substructures most responsible for each outcome, enabling chemists to understand why a molecule behaves as it does and to make informed, targeted decisions during molecular optimization.
Efficient and Scalable
Efficient and Scalable
Efficiency is a core design principle. Our lightweight, scalable models achieve state-of-the-art performance while minimizing computational cost, enabling rapid iteration, fast inference, and seamless deployment through our cloud platform—without sacrificing accuracy or insight.
Explore how our model enables meaningful partnerships.
Partnering